4 minute read.Using the Crossref REST API. Part 12 (with Europe PMC)
As part of our blog series highlighting some of the tools and services that use our API, we asked Michael Parkin—Data Scientist at the European Bioinformatics Institute—a few questions about how Europe PMC uses our metadata where preprints are concerned.
Tell us a bit about Europe PMC
Europe PMC is a knowledgebase for life science research literature and a platform for innovation based on the content, such as text mining. It contains 34.6 million abstracts and 5 million full-text articles. At Europe PMC we support the research community by developing tools for knowledge discovery, linking publications with underlying research data, and building infrastructure to support text and data mining. Our goal is to create a supportive environment around open access content and data, to maximise its reuse.
What problem is your service trying to solve?
Recent years have seen a dramatic increase in the popularity of preprints within life sciences literature. Preprints have been supported by Crossref since November 2016. In response to the rise in popularity, we have started indexing preprints alongside traditional journal publishing within Europe PMC. We expect this will:
- provide another means to access and discover this emergent form of scholarly content
- help explore more transparently the role of preprints in the publishing ecosystem
- support their inclusion in processes such as grant reporting and credit attribution systems
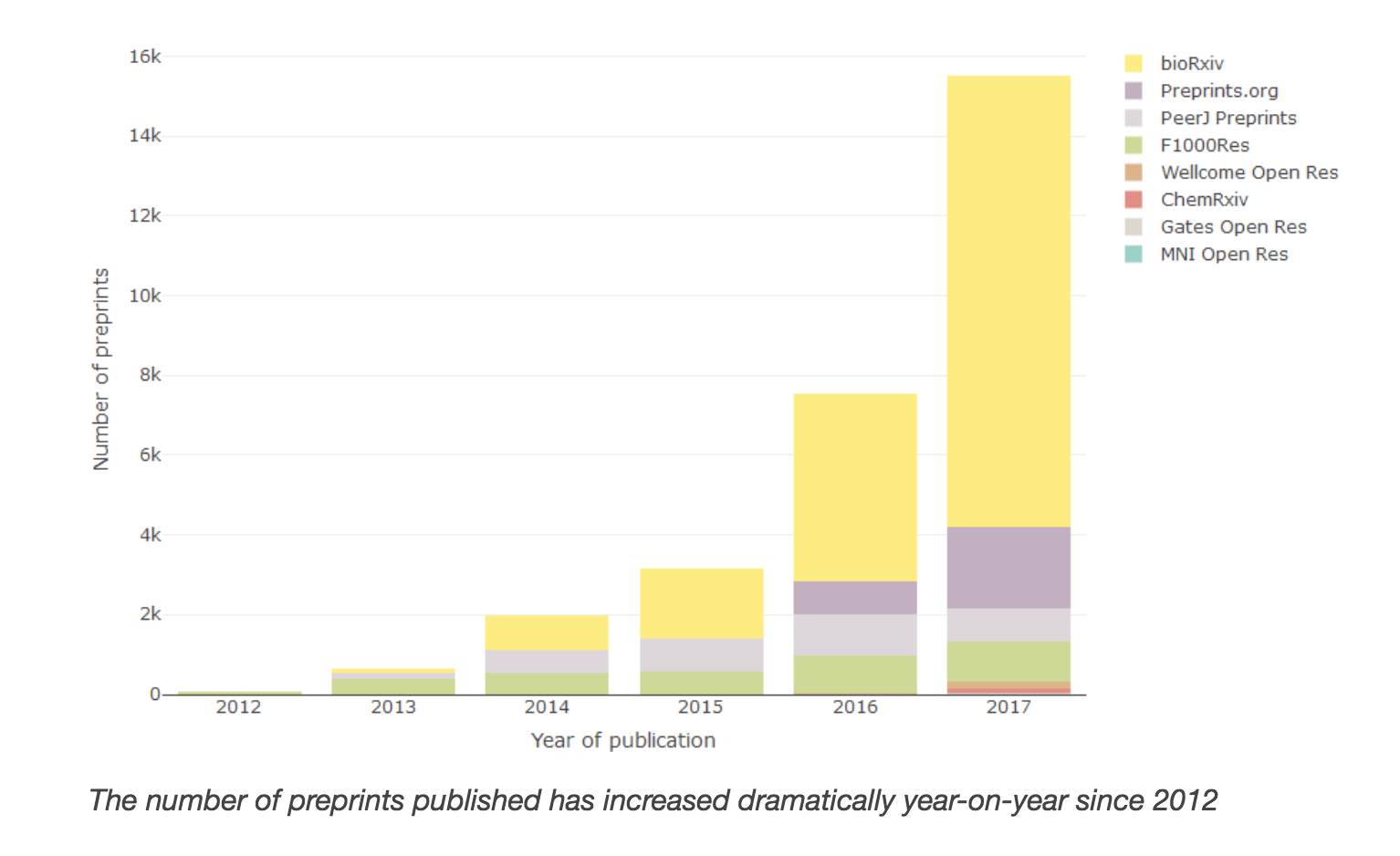
Europe PMC operates an open citation network that uses reference lists from our full-text content, supplemented with metadata supplied by the Crossref OAI-PMH API. The number of citations we retrieve from Crossref increased significantly in 2017 thanks to the efforts of the Initiative for Open Citations (I4OC) in improving awareness about sharing citation data.
Our work to ingest preprints into Europe PMC, however, represents our first use of the Crossref REST API. We make a series of queries for each preprint provider, making use of the “posted-content”, “prefix” and (optionally) “has-abstract” filters. We intend to migrate to using the REST API for the majority of retrievals of Crossref content in due course.
Currently we make use of the following fields:
posted as a publication dateabstractDOIauthor for author given names and surnamestitle as the preprint titleis-preprint-of to establish preprint –> article links
We query the REST API daily making use of the from-index-date filter and cursor pagination to insert new or modify existing records. This means that preprints will be available in Europe PMC within 24 hours of the metadata being sent to Crossref. We store the full REST response in MongoDB, a document-based database. Here are some examples of Crossref API queries used to preprint provider PeerJ Preprints:
calling `https://api.crossref.org/works?filter=type:posted-content,has-abstract:true,from-index-date:2018-07-29,prefix:10.7287&sort=updated&rows=1000&cursor=*`
calling `https://api.crossref.org/works?filter=type:posted-content,has-abstract:true,from-index-date:2018-07-29,prefix:10.7287&sort=updated&rows=1000&cursor=AoN4ldf88uQCe6e1g%2FPkAj8SaHR0cDovL2R4LmRvaS5vcmcvMTAuNzI4Ny9wZWVyai5wcmVwcmludHMuMjcwNjJ2MQ%3D%3D`
Done importing PeerJ Preprints
modified: 2
inserted: 10
From the database we parse out the relevant fields and pass them to our main relational database prior to indexing. This avails the preprint abstracts to all of the value-added services we offer for peer-reviewed abstracts, such as citations, grants, ORCID claiming, text mining, etc. We assign a unique persistent identifier comprising “PPR” followed by a number (1) to each preprint record.
This is displayed on the Europe PMC site as an abstract record, analogous to PubMed records, but with an obvious banner (2) indicating to readers the preprint designation; a tooltip provides further explanation of what a preprint is in comparison to a peer-reviewed article.
Once available on the Europe PMC platform, we then apply downstream processes including:
- providing an Unpaywall link directly to the full-text (3);
- adding a hyperlink to the final published version (if there is one that we can detect) (4);
- incorporating the preprint into our citation network (5);
- adding useful links to e.g. alternative metrics, scientific comments and peer reviews, underlying research data in life science databases (6);
- providing text mined annotations via SciLite (7);
- including funding information (8);
- displaying ORCID claims in the author list (9).

What are the future plans for Europe PMC and preprints?
The inclusion of preprints within Europe PMC is of immediate benefit to researchers who want to explore the very latest research. Moreover we see this as an opportunity for both ourselves and the community to explore how preprints fit into the wider publishing ecosystem; for example to answer questions such as: How often will they be cited? How will they be linked to grant funding and other credit systems? How will they be reused?
What else would you like our API to do?
The REST API and rich metadata model provided by Crossref around preprints are both excellent, but the population of the metadata fields by preprint providers can be limited and/or heterogeneous. The key challenge we see is in encouraging providers to populate the Crossref metadata fields more fully and in a uniform manner.
Thanks to Michael.
If you’d like to share how you use our Metadata APIs please contact the Community team.







